Further feedback very much appreciated, I will update the tutorial with your comments as you send them in. Try to ensure you match up the correct columns. Or it could be that this has something to do with selecting the wrong primary key column in the attribute table (the column that will link the attribute data to the network data you uploaded earlier). The latest generation of Cytoscape (version 3.0 and later) has substantial improvements in function, user interface, and performance relative to previous versions. In that case, try to unselect the relevant columns when uploading them or see if there are any options concerning this in the upload dialog box. Cytoscape provides core functionality to load, visualize, search, filter, and save networks, and hundreds of Apps extend this functionality to address specific research needs. It could also be that the unique ID given to the network table you upload first is the same as the unique ID given to the attribute table you add second. 
In that case, just start with a completely new project. But sometimes this happens when you try to upload the same attributes files twice because it failed the first time. I am on a research trip this month without Cytoscape access. See keep an eye open for that.Ĭoncerning the ‘duplicate column name found’ problem. Such functional categories are typically derived. The latest generation of Cytoscape (version 3.0 and later) has substantial improvements in function, user interface, and performance relative to previous. Given a list of genes resulting from an experiment, enrichment analysis enables to identify functional categories that are over-represented. In April I will make a new tutorial for archaeologists available, using Visone for exploratory network analysis for archaeologists. Enrichment analysis (also known as functional enrichment) is an helpful technique for high-throughput data interpretation. This because the cytoscape updates come quite fast and often are quite significant. It does indeed work with an older version of Cytoscape and I will sadly not update the tutorial anymore (you reminded me that I should put a note about this in the tutorial document). 
It also highlights new features that benefit experienced users.Ĭytoscape interactive network visualization network analysis.Thanks for your kind comment, really delighted that the tutorial is useful for you.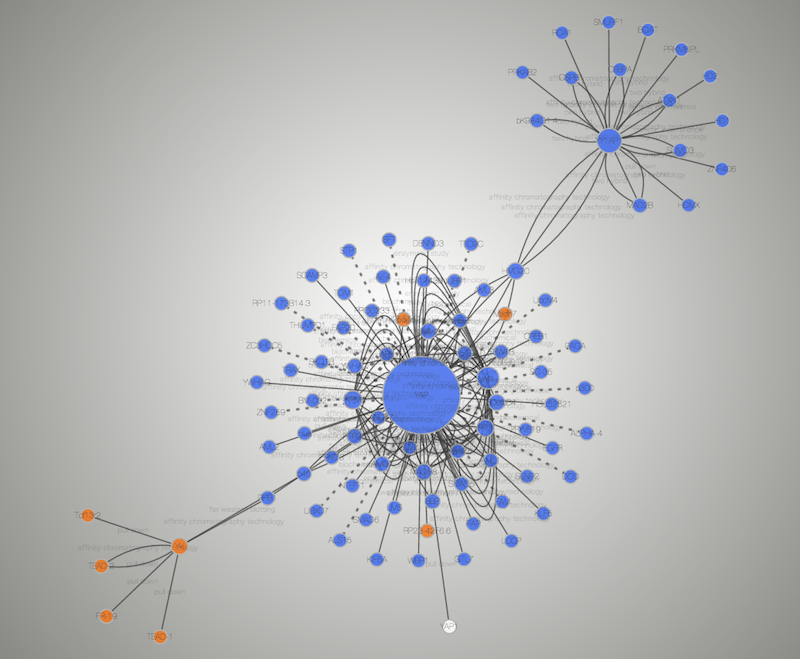 
This protocol aims to jump-start new users with specific protocols for basic Cytoscape functions, such as installing Cytoscape and Cytoscape Apps, loading data, visualizing and navigating the networks, visualizing network associated data (attributes), and identifying clusters. Cytoscape provides core functionality to load, visualize, search, filter, and save networks, and hundreds of Apps extend this functionality to address specific research needs. Example: A vertex represents an airport and stores the 3-letter airport. August 2017 Authors: Pablo Porras EMBL-EBI Abstract and Figures This document is used in presential courses and. It offers researchers a versatile and interactive visualization interface for exploring complex biological interconnections supported by diverse annotation and experimental data, thereby facilitating research tasks such as predicting gene function and constructing pathways. sql is the destination mssql table model. Coloring Nodes by Fold-Change in Cytoscape 3. Introduction to network analysis using Cytoscape and PSICQUIC tutorial. Cytoscape is one of the most popular open-source software tools for the visual exploration of biomedical networks composed of protein, gene, and other types of interactions.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
